Restriction Mapping with CodonCode Aligner
CodonCode Aligner lets you view and print restriction maps for any sequences in your projects. You can choose between single-line, multiple-line, and text only formats:
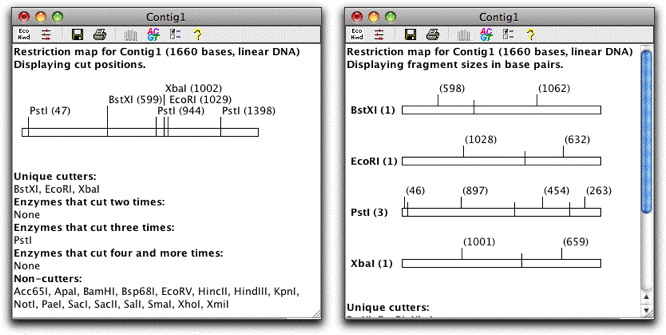
You can select restriction fragments in the map by clicking on the fragment, and then copy the selected bases and paste them into a different program, or create a new text sequence from the selection in CodonCode Aligner.
You can select any set of restriction enzymes to use based on recognition size size, cut result, and how many cut sites are found:
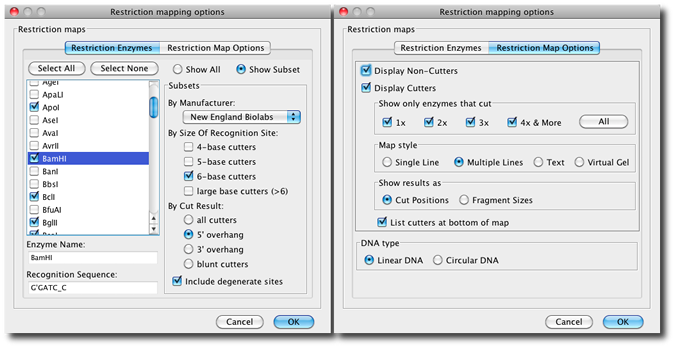
Restriction maps can also be displayed as virtual gels as shown on the right. In the restriction map settings for the virtual gel you can choose the marker for the gel. Another option is to show a lane that includes the fragments for a digest with all enzymes. One lane for each enzyme is always displayed. Mouse overs show the length as well as start and end base for each fragment. As in all other restriction maps in CodonCode Aligner, you can select a fragment by click which will enable copying and highlight the selected sequence in other views: 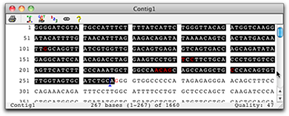 |
 |
To learn more about restriction maps in CodonCode Aligner, look at the restriction mapping tutorial.