Sequence Assembly with CodonCode Aligner
Assemble your sequences quickly and accurately - whether you are building separate contigs for hundreds of different clone or a single contig with thousands of sequences. CodonCode Aligner's sequence assembly features include:
- Fast assembly: Select your sequences, click a toolbar button, and get your contigs in seconds.
- Accurate consensus sequences: CodonCode Aligner uses Phred quality scores to automatically pick the best sequence when building the consensus sequence. This can dramatically reduce the need to edit your sequences.
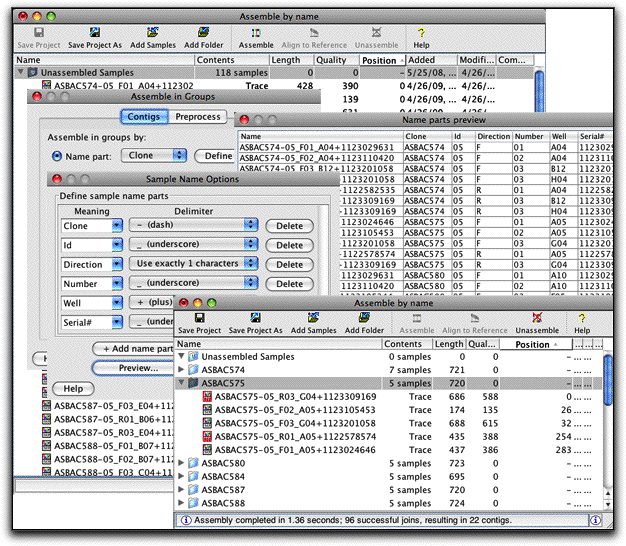
- Build contigs for many clones automatically: Assemble your contigs in groups, based on sample names. Easily customize the way sample names are interpreted for your sequence names.
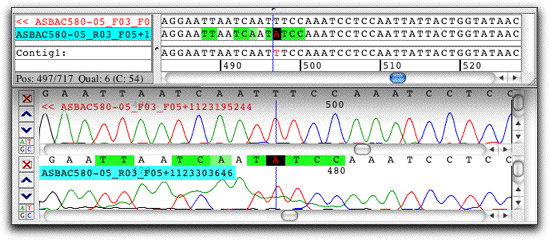
- Compare contigs: Align contigs with MUSCLE or Clustal, and go back to the chromatograms with a simple mouse click.
- One step processing: Have your sequences automatically end clipped, vector screened, and even base called with Phred before assembly.
- Assemble cDNA to genomic sequence.
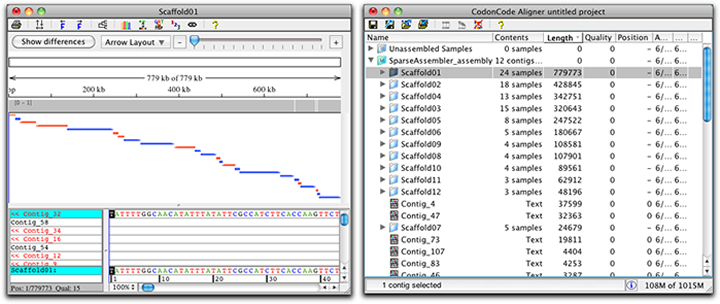
- Assemble and scaffold bacterial genomes. Assemble small NGS data sets like bacterial genomes quickly, and have CodonCode Aligner use information from read pairs to generate scaffolds. Error correct NGS reads before assembly to get larger contigs and scaffolds. Aligner's support for NGS error correction and assembly programs can easily be adapted to allow use of different programs.
- Assemble with Phrap: Use the power of Phrap to assemble projects like whole BAC shotgun assemblies directly from CodonCode Aligner. Then, use CodonCode Aligner's contig editing tools to complete your project.
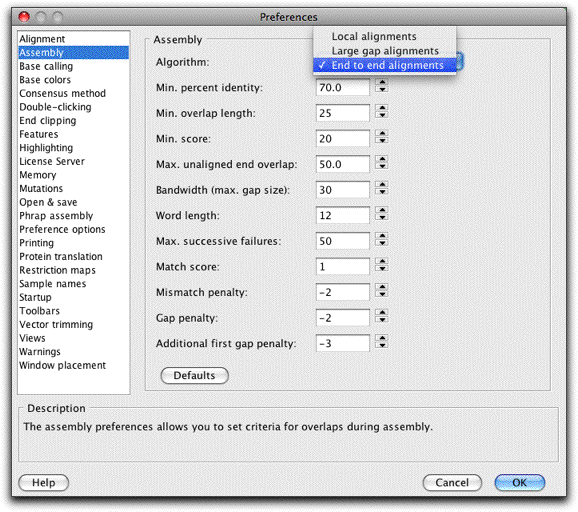
- Control: Set assembly parameters like minimum percent identity, match score,s and gap penalties to control the result of your assemblies.