What's new in CodonCode Aligner 11.0
CodonCode Aligner version 11.0 adds Gibson Assembly, annotate common features, automatic synchroniation of sequence starts, and more.
Please note that CodonCode Aligner 11 requires a new license key. Version 11 is a free upgrade for all users with a current update and support agreement as of July 18, 2023. Other customers can purchase an upgrade to version 11.
Gibson Assembly
Support for virtual Gibson Assembly allows you to simulate your cloning experiment with up to 15 fragments.
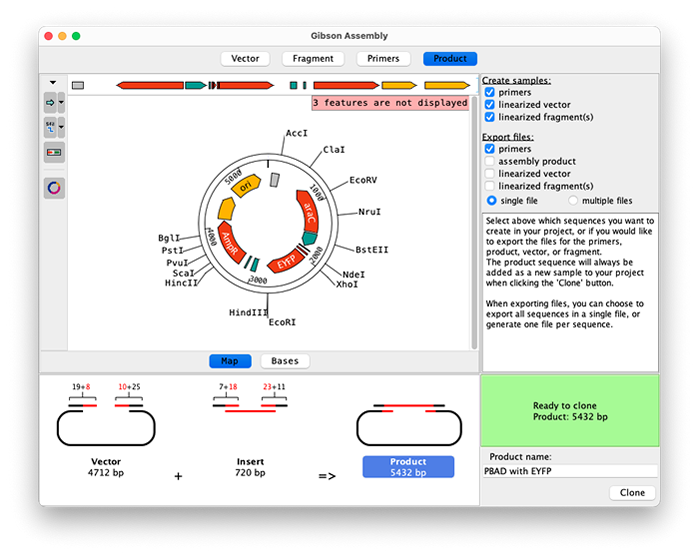
The vector, fragment and product sequences can be displayed as linear or circular maps, or on a base level. Select your region to clone with one of multiple easy options like clicking on a feature, inverting an existing selection, selecting enzymes, entering base numbers, or simply using Shift-click.
The toolbar buttons on the left of the view contain options to change which map, features and enzymes are shown, for example showing only predefined features.
Add up to 15 fragments, and modify, order or delete them as needed.
Automatically pick the best primers depending on your Tm and overlap settings. You can also manually edit primers, for example to add spacers or adjust the reading frame if necessary.
A graphical overview shows your vector, fragment, primers and product.
During your cloning experiment choose to export primers and fragments directly and which sequences to create in your CodonCode Aligner project. |
Find & Annotate Common Features
Better support for features in Aligner 11 includes the option to detect and annotate common features in plasmids. Set the % similarity of features to find and choose which ones are annotated:
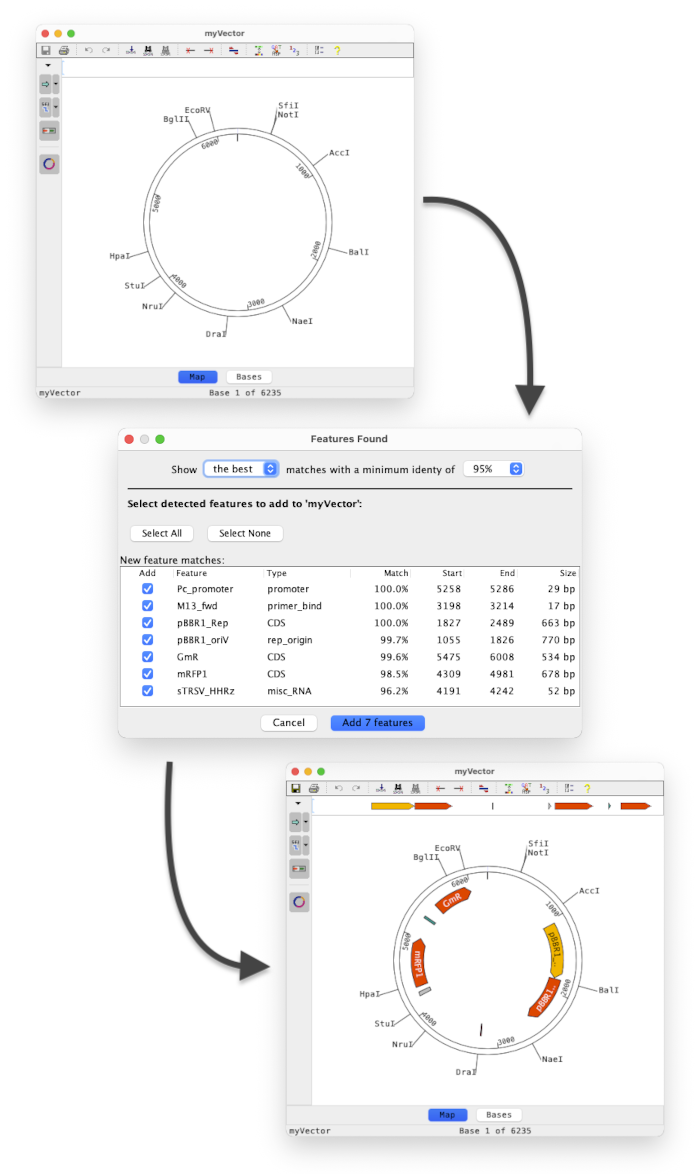
The examples above show a vector before and after finding and annotating common features with at least 95% identity.
|
Synchronize Starts
The function to synchronize the start positions of circular sequences makes it easy to line up and compare similar samples. Here is an example that shows two vectors with some similarities but different start postions before and after synchronizing the starts:

If you prefer you can now also set the start base manually instead of using the automated synchronization feature.
|
More...
- Import features from GFF files
- Set the start in circular sequences
- Better alignments for circular sequences
- Import recent files
|
CodonCode Aligner 11.0 is compatible with Windows 11 and macOS Ventura (13).
Interested? Watch the movie showing the new features of CodonCode Aligner 11.0.1.