Primer Design in CodonCode Aligner
Primer Design in CodonCode Aligner uses Primer3 to find primers, which can then be imported in your Aligner project. Plenty of primer picking parameters like primer characteristics and reaction conditions can be adjusted to create PCR, cloning, and sequencing primers.
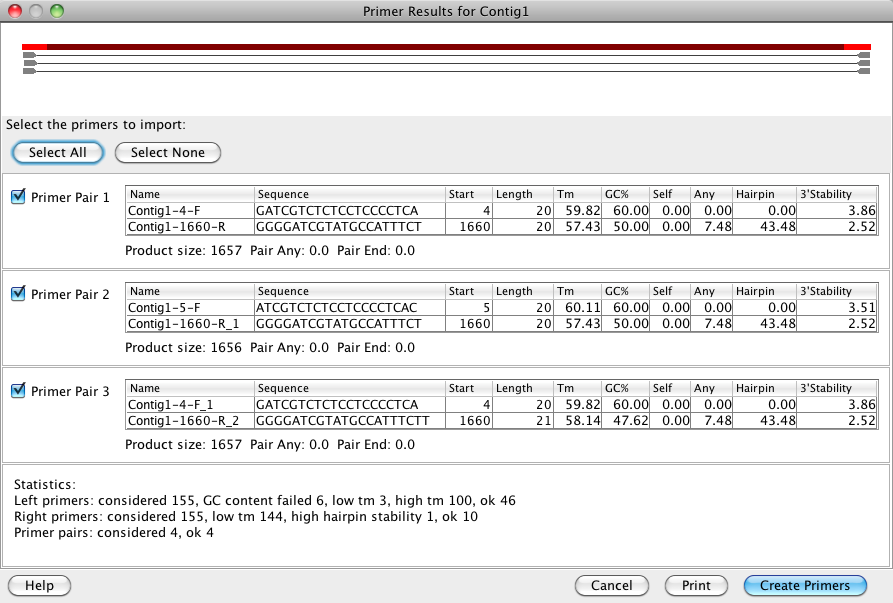
The image above shows the results after finding 3 PCR primer pairs. An overview shows the primer location in relation to the target region. Primers are listed below the graphic along with their characteristics like length, melting temperature, GC content, etc. A statistic about the considered primers and their compatibilty with the chosen settings is included at the bottom. These primer results can be printed as seen on the screen. Primers can be selected and created in your Aligner project for further analysis and for ordering.
Imported primers are created in a separate folder and are highlighted in your target sequence. Primer details can be easily accessed for each primer, or all primers can be viewed together using the features table.
Primers can be exported in a csv file format for ordering.
To learn more about primer design in CodonCode Aligner, have a look at the primer design tutorial.